Understanding how the diversity of life on earth arose and is maintained is the foundational question in evolutionary biology. Our lab studies the processes of adaptation and speciation from a genetic perspective. We are particularly interested in connecting genetic processes to evolution in nature. Current work in our lab focuses on understanding the mechanisms of adaptation, genome evolution, and the consequences of hybridization between species.
What is the genetic architecture of reproductive isolation between hybridizing species?
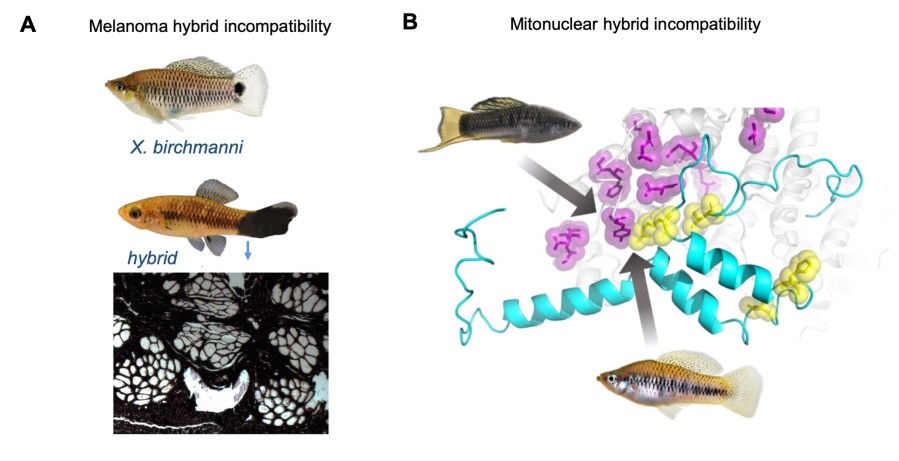
Biologists have long appreciated the fact that when species hybridize, hybrids often suffer reduced viability or fertility, presumably due to genes that have functionally diverged between the parental species and no longer interact properly in hybrids. Despite a century of work documenting this phenomenon, we know very little about which genetic interactions cause hybrid incompatibilities and the evolutionary and molecular paths through which they arise. This is especially true for species that naturally hybridize, many of which have historically been difficult to study in the lab. We use naturally hybridizing species of swordtail fish as a powerful model system to study the genetic basis of hybrid incompatibilities and uncover their evolutionary history. Current work in the lab is focused on the role of coevolution in regulatory networks and protein complexes in driving hybrid incompatibilities.
How does selection shape ancestry in the genome?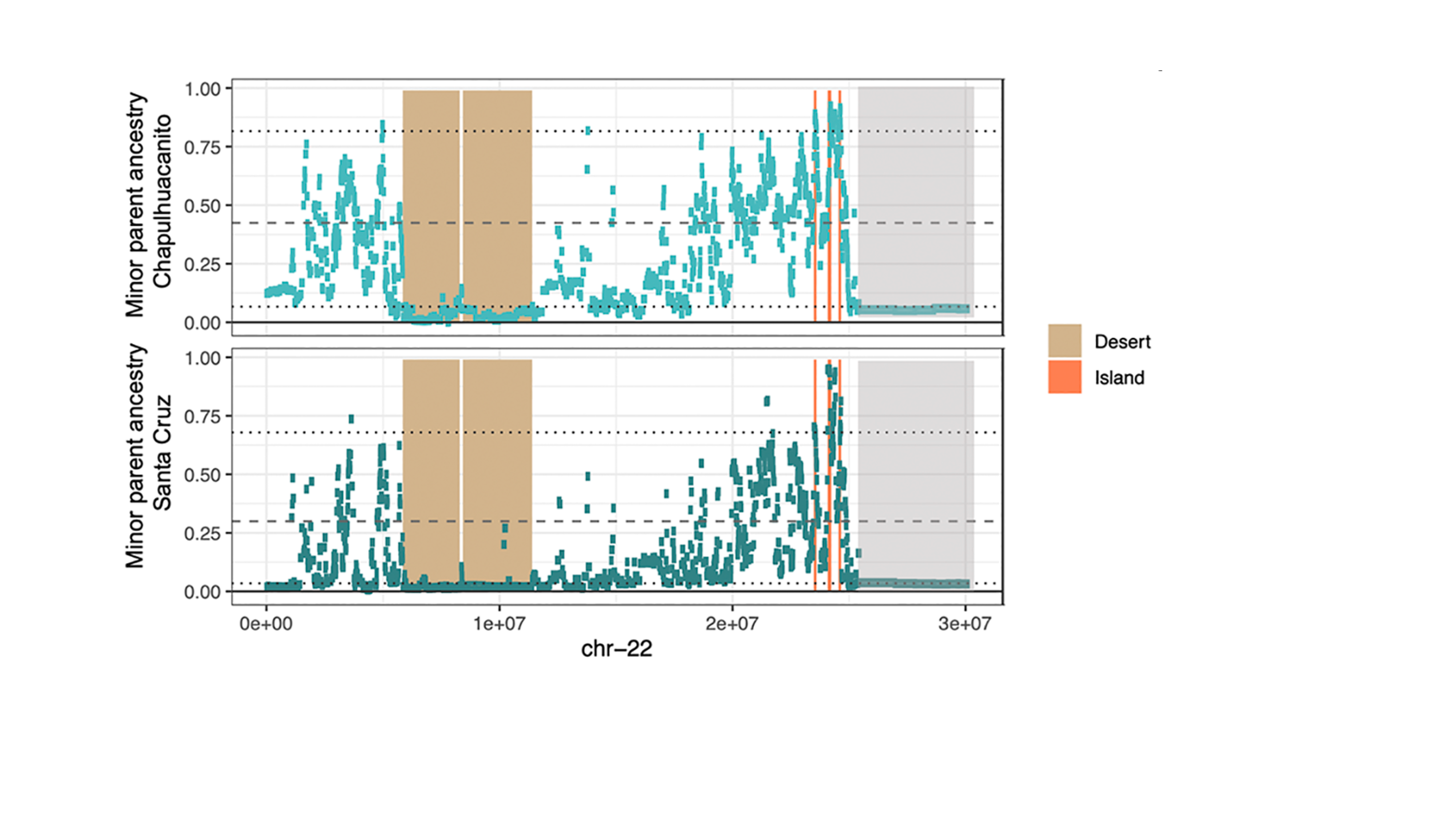
Many species derive large proportions of their genome from hybridization. In some human populations, more than 2% of the genome is derived from admixture with our extinct relatives, while in some swordtail fish species, as much as 30% of the genome is derived from hybridization. However, there is striking variation in the retention of hybridization-derived haplotypes along the genome. Work in out lab seeks to understand how the genome evolves and stabilizes after hybridization, how repeatable these outcomes are, and how basic genetic processes like recombination influence this process.
What is the genetic basis of adaptive differences between species?

Adaptation to the ecological and social environment has a fundamental impact on survival and reproduction. Despite decades of intensive study, we still have an incomplete understanding of how this occurs on the molecular level in natural populations. Our lab is interested in characterizing the genetic basis of such differences between species and uncovering how these loci impact hybridization between species. Current project are focused on the genetic basis of adaptation to different thermal environments and the drivers of sexually dimorphic traits within and between species.
What is the link between adaptation and mechanisms of genome evolution?

A new era of long-read genome sequencing is highlighting exceptional variation in rates of evolution across the genome. Certain regions of the genome because of their structural, repetitive, or epigenetic properties, are especially prone to mutation within species. Our lab has been investigating the role of such regions in evolution of both trait differences and reproductive barriers between species.
CICHAZ field station – Calnali, Hidalgo, Mexico
We work with a fantastic field station in the state of Hidalgo, Mexico. For more information about working at CICHAZ, see the CICHAZ homepage



